Sequence search¶
Landing page¶
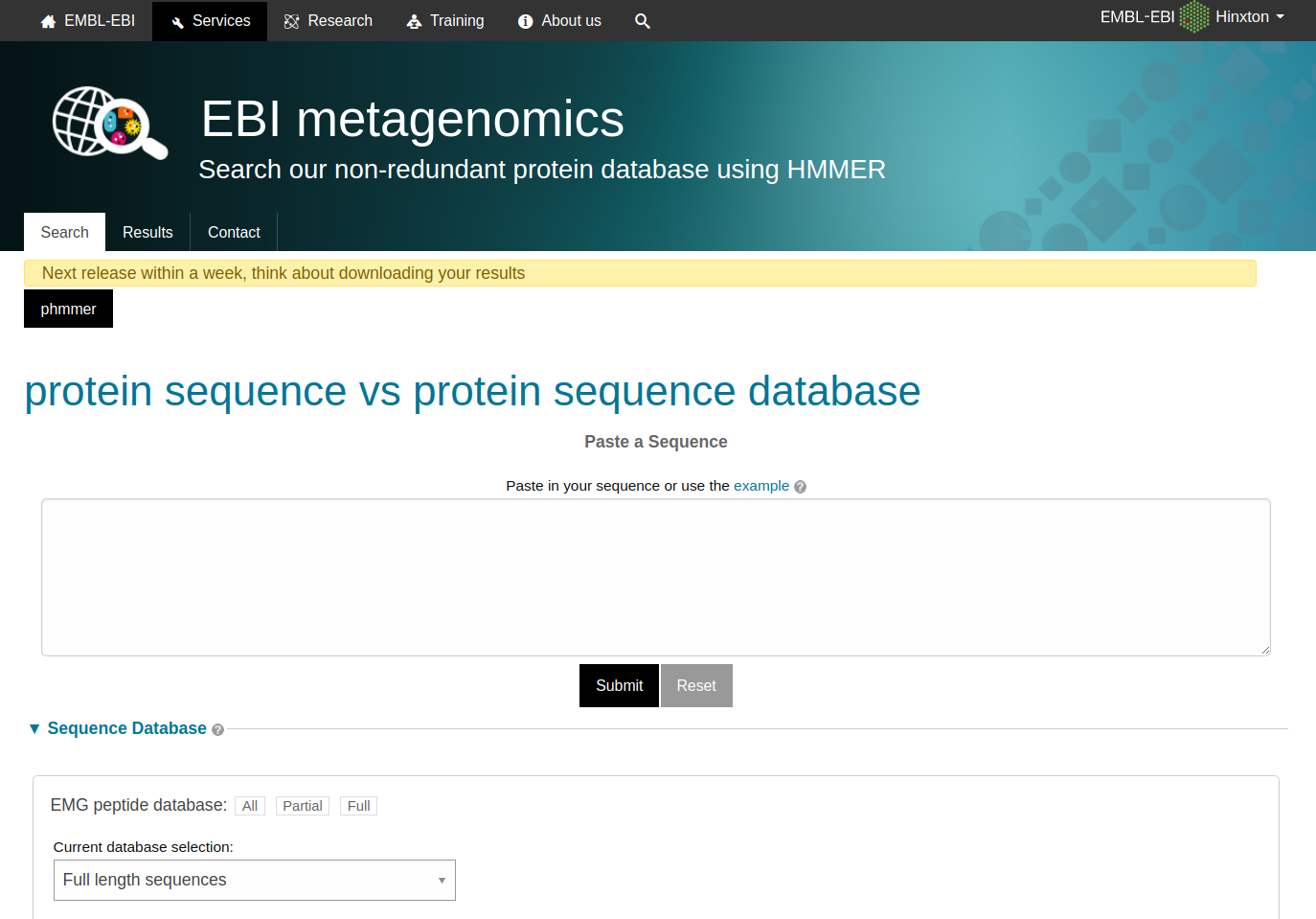
The sequence search (accessed by following the ‘Sequence search’ link from menu bar) provides a search against a catalogue of predicted peptides.
Figure 1. The landing page of the sequence search tool
These sequences comprise a non-redundant set of proteins predicted from contigs that have been assembled from sequencing runs. The HMMER search engine has been adapted to provide fast searches against this database. The results can be linked back to the sample and run from which the peptide was derived and also to sequences with an exact match in the UniProt database.
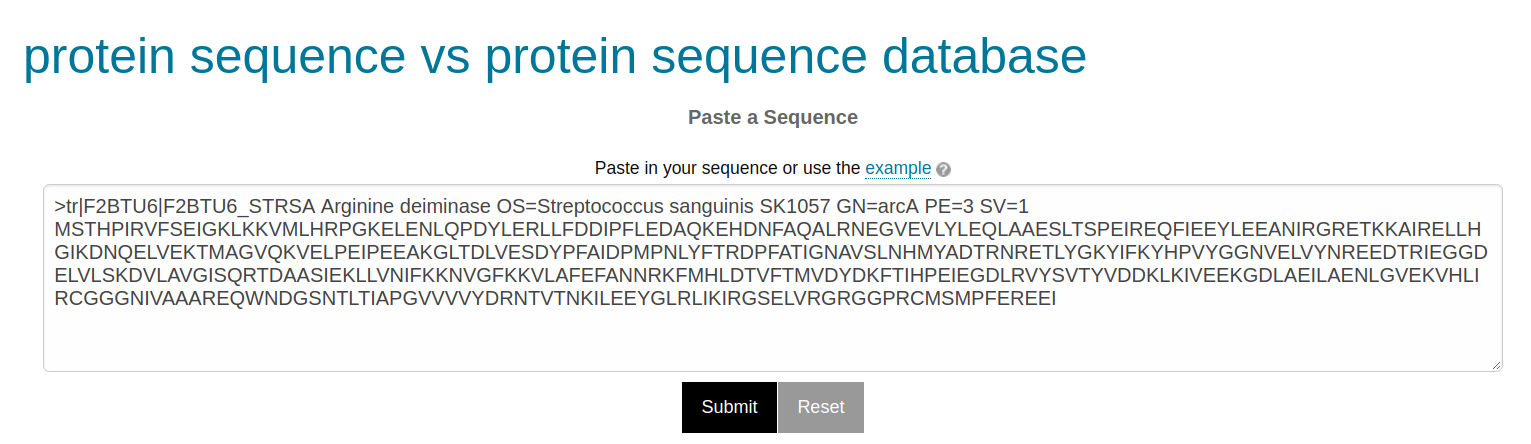
The search takes a FASTA-formatted amino acid sequence.
Figure 2. Example of a well-formatted input sequence
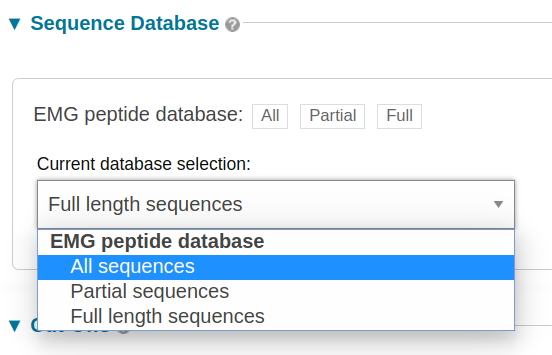
You can search against all of the sequences in the database (‘All’), or restrict your search to full length sequences or partial sequences only (see Partial and full length peptides).
Figure 3. How to select the peptide database to search against
Result page¶
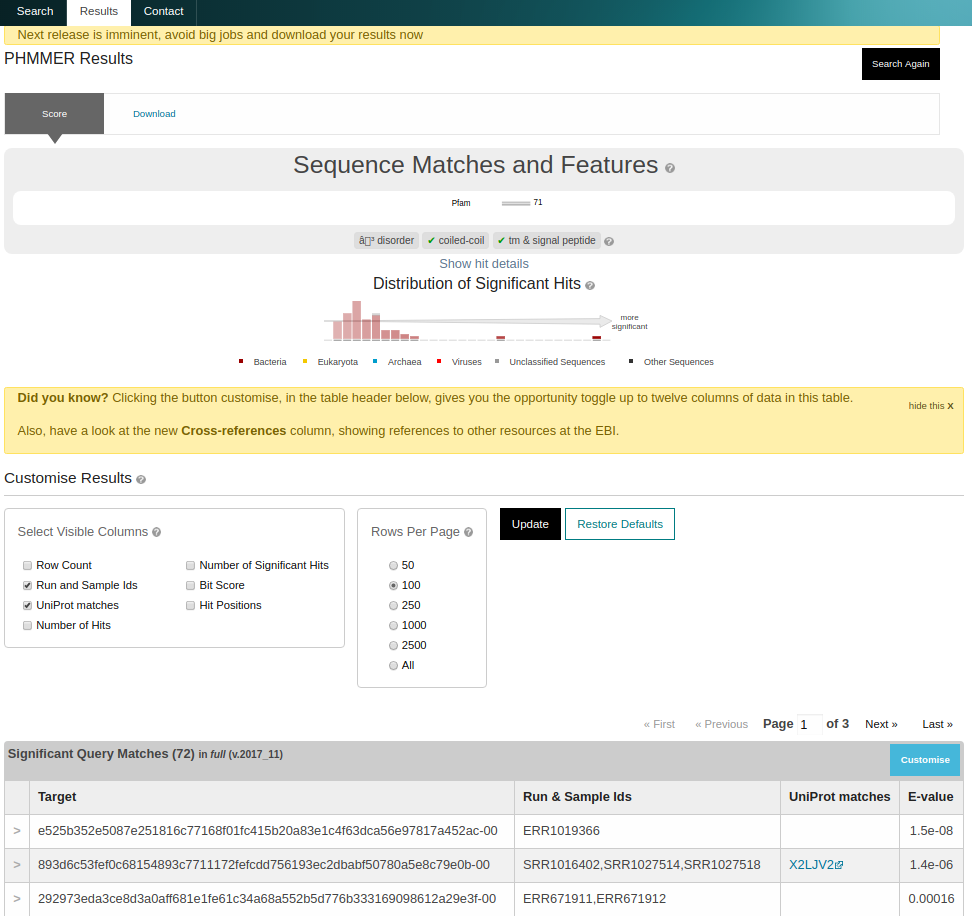
On completion, a list of matching sequences is shown in order of E-value significance. Since identical peptides could be derived from different samples and runs, we use a unique hash sum (SHA256) as the sequence identifier. The mapping to UniProt identifiers and EBI Metagenomics run/sample accessions can be switched on by selecting ‘Customise’ on the results page and checking the appropriate boxes.
Figure 4. Different features on the result page after triggering a sequence search
At this time, it is not possible to link directly to the matching sequence from the results table. However, in the download tab, the ‘Full length FASTA’ link will provide all the matching sequences. Alternatively, the sequences are available on our FTP server (ftp://ftp.ebi.ac.uk/pub/databases/metagenomics/peptide_database).
Build process¶
The database is updated periodically and is created as follows:
- Short reads from runs are assembled into contigs using metaSPAdes
- Contigs are filtered by length (minimum 500 base pairs)
- Peptides are predicted using a combined gene caller (Prodigal and FragGeneScan)
- Resulting peptides are made non-redundant to produce a set of unique sequences
- Sequences are mapped back to EBI Metagenomics run and sample accessions
- Sequences are compared to UniProt and accessions for matching sequences are mapped
- Domain architectures are identified using the Pfam database
Each update (versioned using the release year/month) is cumulative and uses all predicted peptides available at that time.
Partial and full length peptides¶
In common with some other protein coding sequence predictors, Prodigal provides an indication as to whether a gene is full length or extends beyond the contig. To indicate this, the sequence ID has two digits appended (one for each end of the sequence), each of which is either 0 (the gene is encoded within the contig) or 1 (it extends beyond). Thus a full length sequence is suffixed with ‘-00’ and a partial with ‘-11’. The notation ‘-10’ or ‘-01’ is used for the cases where the gene is truncated at only one end. Based on this information, three peptide sequence sets are available for searching: peptides derived from full length genes, peptides derived from partial genes, and all peptides.
>seq_1 # 3 # 371 # 1 # ID=1_1;partial=10;start_type=Edge;rbs_motif=None;rbs_spacer=None;gc_cont=0.501
SEGCEYLAAYLDKRIASGETINESSAVMTLSQGYLMKGRNKDAGKKFITTPAITKEIREA
QT
>seq_2 # 4738 # 5193 # -1 # ID=1_9;partial=00;start_type=ATG;rbs_motif=None;rbs_spacer=None;gc_cont=0.568
MSAYWYAVIWGGSFGAVLAAAGPRFRKAIPAIRGRMKNSIKWSTSAKAINGISWAGPFAA
QT
>seq_3 # 7546 # 8232 # -1 # ID=1_11;partial=00;start_type=TTG;rbs_motif=GGAG/GAGG;rbs_spacer=5-10bp;gc_cont=0.541
MKKKVLSIQNIACETLGTLEGMFRKDGLEVENVSAQEGGIPIKSSEYSAVVVLGGPMAVY
QT
>seq_4 # 32 # 103 # -1 # ID=37115_1;partial=01;start_type=Edge;rbs_motif=None;rbs_spacer=None;gc_cont=0.542
WILDGIDIDAMIRHPVRQYQIAG
Availability¶
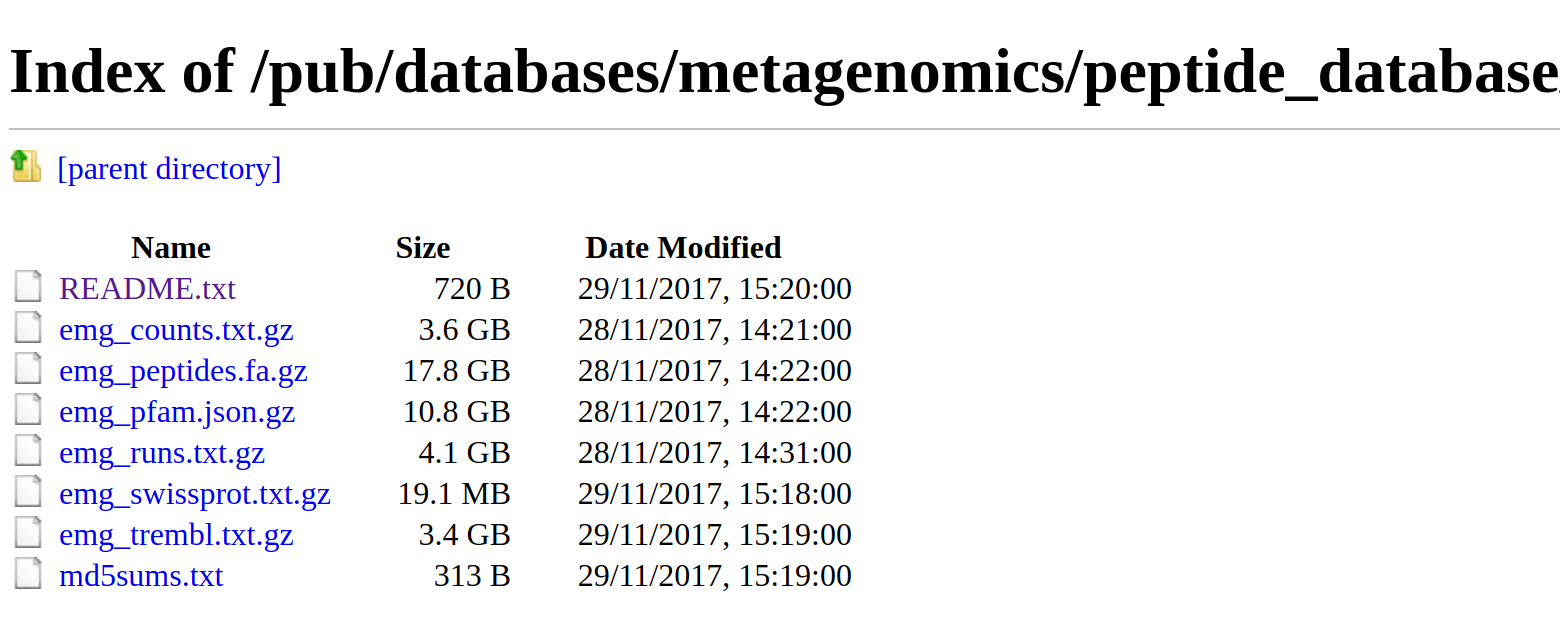
As well as searches via a web server, we provide all data for download from our FTP server (ftp://ftp.ebi.ac.uk/pub/databases/metagenomics/peptide_database). This includes the sequence database, run, sample, UniProtKB/SwissProt and UniProtKB/TrEMBL mappings, Pfam architectures, and counts of the number of times each sequences was observed in the database as a whole.
Figure 5. List of available files on the FTP server
Further information¶
Full documentation regarding the HMMER webserver is available. Note that some of the documented features (such as the taxonomy view) are not relevant to the peptide search and are therefore disabled. If there are additional features or feedback on this search service, please get in contact with us.